This package contains methods for clustering, pairing and testing single cell RepSeq data, especially as generated by 10X genomics VDJ solution.
Installation
Install with
install.packages('BiocManager') # if you don't have it yet
BiocManager::version() # Check that Bioconductor version >= 3.12
BiocManager::valid() # And it's in a sane state.
BiocManager::install('CellaRepertorium') # installFor the development version, install via
devtools::install_github('amcdavid/CellaRepertorium').
Data requirements and package structure
The fundamental unit this package operates on is the
contig, which is a section of contiguously stitched
reads from a single cell. Each contig belongs to one
(and only one) cell, however, cells generate multiple contigs.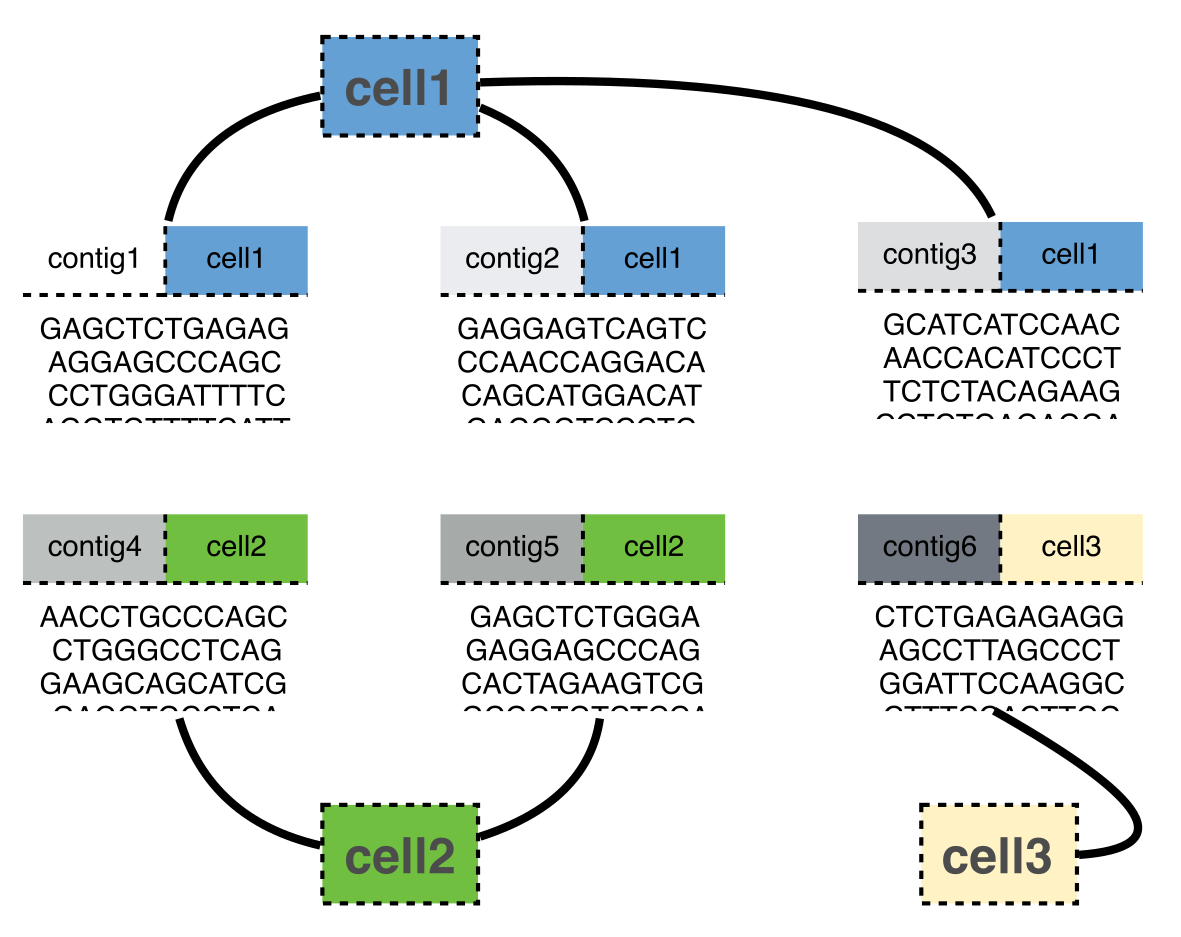
Contigs can also belong to a cluster. Because of
these two many-to-one mappings, these data can be thought as a series of
ragged arrays. The links between them mean they are relational data. A
ContigCellDB() object wraps each of these objects as a
sequence of three data.frames
(dplyr::tibble(), actually). ContigCellDB()
also tracks columns (the primary keys) that uniquely identify each row
in each of these tables. The contig_tbl is the
tibble containing contigs, the
cell_tbl contains the cells, and the
cluster_tbl contains the clusters.
The contig_pk, cell_pk and
cluster_pk specify the columns that identify a contig, cell
and cluster, respectively. These will serve as foreign keys that link
the three tables together. The tables are kept in sync so that
subsetting the contigs will subset the cells, and clusters, and
vice-versa.
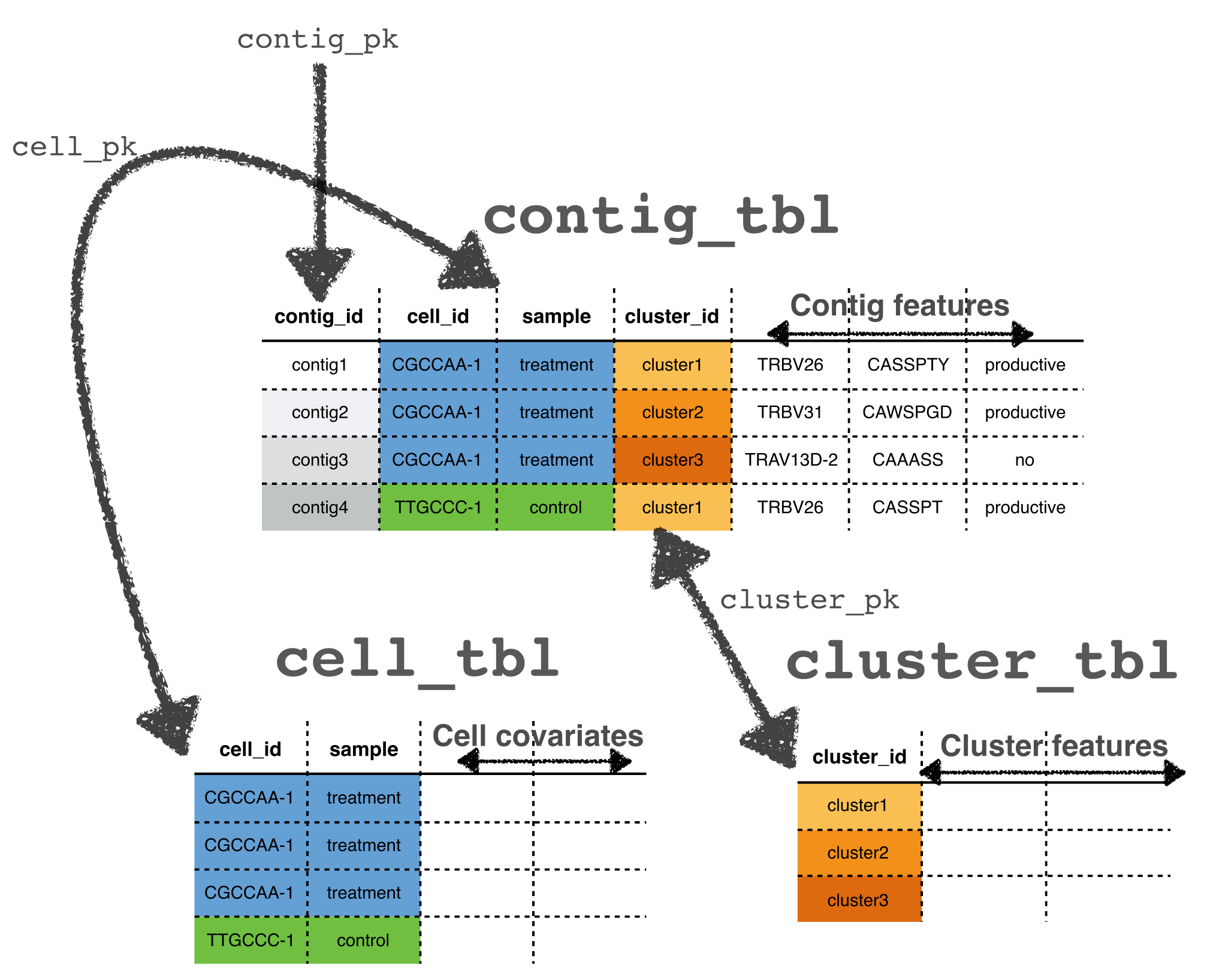
Of course, each of these tables can contain many other columns that will serve as covariates for various analyses, such as the CDR3 sequence, or the identity of the V, D and J regions. Various derived quantities that describe cells and clusters can also be calculated, and added to these tables, such as the medoid of a cluster – a contig that minimizes the average distance to all other clusters.
Some functions of interest
Mainly, this package seeks to enforce proper schema of single cell repertoire data and stay out the user’s way for various summaries they might conduct.
However, there are a variety of specialized functions, as well:
-
cdhit_ccdb(): An R interface to CDhit, which was originally ported by Thomas Lin Pedersen. -
fine_clustering(): clustering CDR3 by edit distances (possibly using empirical amino acid substitution matrices) -
canonicalize_cell(): Return a single contig for each cell, e.g., for combining VDJ information with 5’-based single cell expression -
ccdb_join(): join accdbobject from this package to aSingleCellExperimentobject, by droplet barcode. -
cluster_permute_test(): permutation tests of cluster statistics -
cluster_logistic_test(): logistic regression tests for overrepresentation of clusters among cells -
pairing_tables(): Generate pairings of contigs within each cell in a way that they can be plotted
Interfacing related packages for clonal analyses
- To combine repertoire information with expression of endogenous
mRNAs, this package has been used with
SingleCellExperiment::SingleCellExperiment()and Seurat after generating cell canonicalizations. - Many tools from the Immcantation
suite can work directly on
ContigCellDB()objects.